|
Once you have an aligned movie you can draw the bleaching ROI and run Plot Z-axis profile (under Images › Stacks). Of course, this plugin does not always work - if your sample moved a lot or if the signal does not allow for efficient comparison, the resulting movie will be pointless. This way you are not playing around with the fluorescence intensity values and you do not have to manually adjusting your ROI frame by frame. You can minimize this by using the Linear Stack Alignment with SIFT plugin designed by Stephan Saalfeld which basically compares a frame with its previous one and tries to compensate eventual movement by rigidly moving the whole image so that they better overlap. If it has moved during image acquisition, so will have your bleaching ROI. Once you have done this, you should carefully look at your specimen. For this I use the Concatenator plugin for ImageJ which can be found here (). 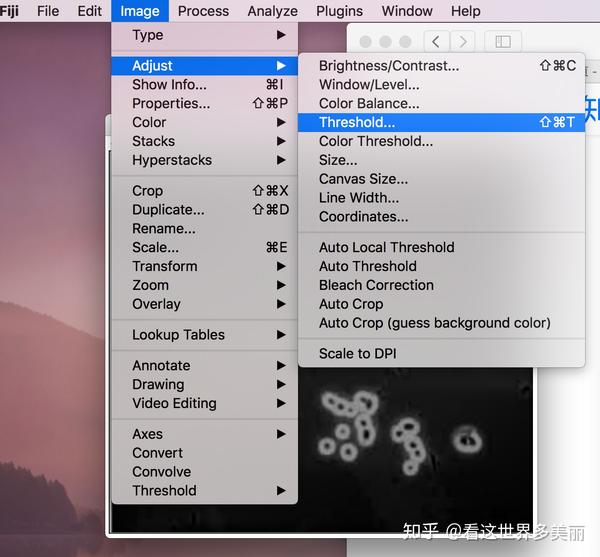
The next step will be to unite the 4 movies in a single one. Load the 4 different time series in Fiji. It is now time to analyze the results… Analysing the data Once everything is done you should have 4 different time series and 205 frames in total. All this can be set up automatically by using the Visual Macro option in the DuoScan confocal system. And finally you can take frames every 30s for another 10minutes - slow acquisition postbleach. After this minute of intense imaging you can delay your frame rate to a medium level (one frame every 5s for 10 minutes) - medium acquisition postbleach. Immediately after the bleach you take 60 frames every second - this will be your fast acquisition postbleach time series (1m). After this initial time series, you bleach your sample using the conditions previously described. The first 5 frames are taken every 5s without any bleach - this will be your prebleach conditions (25s). However there are different acquisition rates during this time. In my particular case, every experiment lasts for 21m25s. Now you have to establish the duration of the experiment - this depends on the kinetics of recovery of every protein so optimization is advised. I usually take 512x512 frames with a zoom of 3 and time of acquisition around 1s. Therefore you have to get quick time frames with a good enough resolution for your further analysis and quantification. This is crucial for the aspect of your fitting curve. Since you are interested in obtaining a good description of the fluorescence recovery it is important to get the maximum number of time points immediately after the bleach. The number of iterations used is 7 - this is probably one of the most important aspects of the experiment: depending on the protein, model organism and size of the region of interest, this number should be increased (this might result in ablation) or decreased (which can cause no bleach at all) according to your needs. The bleaching conditions are as follow: 488nm laser at 100% and 405nm laser at 100% also. In the channel view, I take sections of 2μm (this can be adjusted by the pinhole) and usually slide the gain to around 750. For the acquisition itself, I set the laser at 10%. I set up the Argon laser output power to 45% so that the current through it is around 6A. I normally use GFP tagged proteins, so the acquisition laser used is the Argon 488nm and for the bleaching I use the 405nm diode. Optimization of the bleaching and acquisition parameters while using other model organisms is strongly recommended. The following tutorial has been optimized to Drosophila embryos using the DuoScan confocal system in the MPI-CBG. 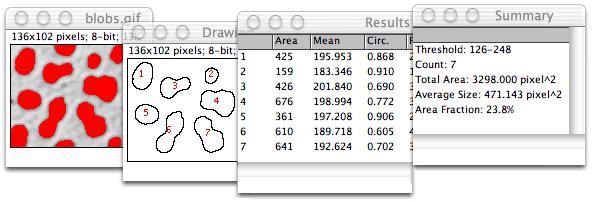
Fluorescence Recovery After Photobleaching in Drosophila embryos At the confocal Comments on the content of this tutorial are welcome. 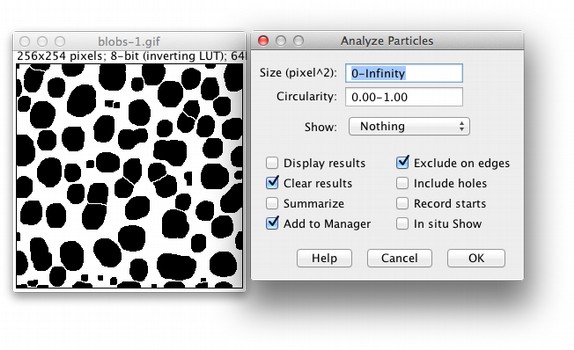
It relates how to correct for the drift of your biological samples during long-term timelapse imaging for subsequent analysis using Fiji. This tutorial is brought to you by Joao Firmino, Knust lab, MPI-CBG. If you’d like to help, check out the how to help guide! The content of this page has not been vetted since shifting away from MediaWiki.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
